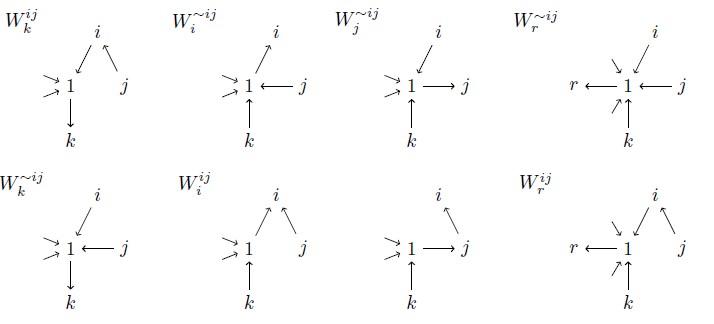
Several possible configurations of a star network, with 1 as a hub at the center. This kind of network can represent travel between a city and the suburbs. Courtesy of Stephen Kirkland, Zhisheng Shuai, P. van den Driessche, and Xueying Wang.
The COVID-19 pandemic has made clear the importance of understanding precisely how diseases spread throughout networks of transportation. However, rigorously determining the connection between disease risk and changing networks--which either humans or the environment may alter--is challenging due to the complexity of these systems.
In a paper published in the SIAM Journal on Applied Mathematics, Stephen Kirkland (University of Manitoba), Zhisheng Shuai (University of Central Florida), P. van den Driessche (University of Victoria), and Xueying Wang (Washington State University) study the way in which changes in a network of multiple interconnected communities impact the ensuing spread of disease. The four researchers were hosted as a Structured Quartet Research Ensemble by the American Institute of Mathematics.
A common mathematical model uses several interconnected patches to represent separate geographical regions that are connected by transportation networks. Diseases are often transmitted along these kinds of networks via insects like mosquitoes and ticks, which may hitch rides on people or goods. Pathogenic microorganisms, such as bacteria and protozoa, can also spread diseases through river networks.
"This disease transmission could increase due to flooding, which could possibly create a new shortcut," Shuai said. "How would disease dynamics then change in response to this change in the network?"
To answer this question, the researchers sought to measure the basic reproduction number R0 of the network as a whole. R0 determines a disease's invasibility -- if it is greater than 1, the number of infections will most likely grow; if it is less than 1, the disease will eventually die out.
"When the dispersal between patches is faster than the dynamics of the disease or population, it turns out that the network reproduction number R0 can be approximated as a weighted average of the individual patch reproduction numbers," Wang said.
For example, if pathogens in a river are infecting people with cholera and the water is moving faster than the pathogens decay, one can approximate R0 for the entire river network as a combination of the basic reproduction numbers for each separate community along the river. This is important because the value of R0 can guide disease control strategies -- though the information it provides is limited, and it cannot predict the actual size of an outbreak.
The authors developed new techniques based on several areas of applied mathematics to determine how R0 changes when the structure of a network is altered. Their mathematical approach enabled analysis on two different types of model networks: a star network, which contains several branches that stem from a central hub, and a path network, which consists of several communities that are located sequentially along a track.
"A star network can represent human transport between one hub--like a large city--and several leaves, which would represent small cities or suburbs," Wang said. "A path network can represent communities along a river or stream." These frameworks are also flexible -- for example, the star network is useful for modeling multiple possible scenarios.
"In the star network, we can think of a central water source--the center of the star--with several communities supplied by that source," van den Driessche said.
It is possible to add an arc to the path network that bypasses several locations along the river, which could represent a major flood. If a new arc appears that connects a downstream patch to an upstream patch, for instance, the team's model indicated that the disease transmission risk decreased at downstream locations and increased at upstream locations. The model also incorporated a certain "hot spot" along the river at which the disease transmission rate is higher; the bypass could potentially avoid this location. In an example scenario of a path network with five patches numbered 1 (most upstream) through 5 (most downstream) in which there is a bypass from patch 2 to 4, hot spots at different locations produce different effects. When patch 3 is the hot spot, there is no change in R0 for the whole river network; a hot spot at patch 1 or 2 leads to a decrease in R0, while a hot spot at patch 4 or 5 leads to an increase in R0.
The authors utilized their results to explore possible strategies for controlling disease outbreaks by introducing new connections on a network or changing the strength of existing connections. "Our findings from both the star and the path networks highlight that the placement of the hot spot and the connections among patches are crucial in determining the optimal strategy for reducing the risk of an infection," Wang said. The researchers' techniques quantified the effectiveness of different approaches in controlling invisibility and found the mathematical conditions under which it is best to change the amount of movement between certain locations.
The insights from this study could be helpful in forming future disease intervention strategies. "In some practical settings, we may not have much control over the level of invisibility in the individual patches, but we may have better control over the structure of the network connecting those patches -- for example, in a network of airports," Kirkland said. "The insights gained from our research may inform network-based strategies to control the invisibility of disease."
Source article: Kirkland, S., Shuai, Z., van den Driessche, P., & Wang, X. (2021). Impact of varying community networks on disease invasion. SIAM J. Appl. Math.
Society for Industrial and Applied Mathematics