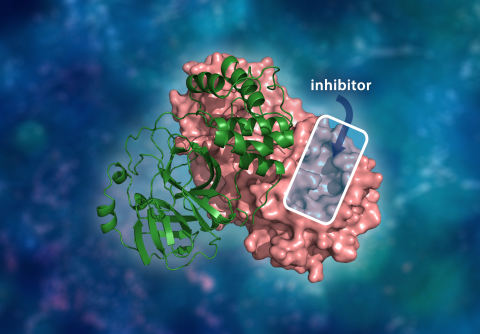
The three-dimensional structure of the SARS-CoV-2 main protease. The molecular dimer complex consists of a single monomer (green) bound to another monomer (pink) where a drug inhibitor would bind to a catalytic site cavity to prevent the virus from replicating. Courtesy of ORNL/Andrey Kovalevsky, Jill Hemman
As the novel coronavirus continues to spread, researchers are searching for novel ways to stop it. But for two scientists, looking to the future means drawing inspiration from the past.
In January of 2020, Andrey Kovalevsky and Daniel Kneller, researchers at the Department of Energy (DOE)'s Oak Ridge National Laboratory (ORNL), were preparing to use neutrons to study the relationship between a certain HIV protease—a protein enzyme that allows the virus to replicate itself within the human body—and a class of anti-retroviral drugs known as HIV protease inhibitors. Some types of HIV build resistance to these drugs. The researchers’ goal was to gain a better understanding of how protease variations work, to aid the development of cutting-edge treatments to overpower even the toughest resistant strains of HIV.
When the team began their work, little did they know that, coincidentally, their efforts to study HIV would quickly put them on a new path to tackling COVID-19, the pandemic that now has the world in its grip.
As it turns out, the protease enzymatic activity that enables HIV to reproduce—the very mechanism Kovalevsky’s team was gearing up to investigate with neutrons—is the same replication mechanism employed by SARS-CoV-2, the virus that causes the disease COVID-19.
Now, the team has shifted the focus of the experimental approach they intended to use to study HIV to combat the new global threat.
Kovalevsky has been studying HIV for 15 years. As a neutron crystallographer, he studies small crystallized samples of biological matter by bombarding them with neutrons. The neutron scattering technique is highly effective in revealing how a sample’s atomic structure is arranged and how its atoms are behaving. Depending on the aim, insights gleaned can offer guidance on how to either improve or even suppress certain properties of a biological material.
Neutrons are an ideal tool for studying biological structures and behaviors because of their acute sensitivity to light elements such as hydrogen and their ability to probe such materials without damaging them.
In 2019, Kovalevsky set out to study HIV in a way that had never been done before. Using inelastic neutron scattering would allow him to collect data on the dynamics, or the motions, of an HIV protease, which would add to the neutron diffraction data he’d been collecting for years. Having both the structural and behavioral—or dynamical—information would provide a more complete picture of how the virus works and, in turn, could lead to new advances in treatments.
After using the VISION spectrometer at ORNL’s Spallation Neutron Source (SNS)—a neutron scattering instrument that reveals the motions of atoms based on their vibrations—Kovalevsky realized he needed help in analyzing the data.
“Daniel brings in expertise in viral protease research,” explained Kovalevsky on recruiting Kneller. “He knows how to work with the proteins in the lab. He knows all the lab techniques in terms of protein production, purification, crystallization, crystallographic data collection, and analysis to obtain insights into drug design.”
It took about 8 months to hire Daniel after an extensive search, Kovalevsky says. Kneller—who specializes in studying HIV protease using crystallography—joined Kovalevsky’s team in January of 2020 to help with the experimental and computational work on the HIV protease.
But just as the team was ready to dive in, COVID-19 had gone global, and the research hit a hard stop.
In March, staff in ORNL’s Neutron Sciences developed a plan to study key components of COVID-19 by assembling research teams and reprioritizing the operating schedules of essential instruments at the two neutron scattering facilities at ORNL, SNS and the High Flux Isotope Reactor (HFIR).
Having already laid the groundwork to study protease, Kovalevsky and Kneller promptly pivoted from HIV to the novel coronavirus. Specifically, they are currently focused on the main protease of SARS-CoV-2, the virus that causes the COVID-19 disease.
“The SARS-CoV-2 protease is an enzyme that cuts proteins that enable the virus to reproduce. Understanding how the protease is assembled and how it functions is a critical first step to finding effective drug inhibitors to block the virus’s replication mechanism,” said Kovalevsky. “Similar to the HIV protease, the main protease from the SARS-CoV-2 virus is one of the most attractive drug targets right now for designing specific inhibitors.”
As with the original plan of the HIV work, the team is preparing to use instruments at SNS and HFIR to gain fundamental insights into how the atoms in the protease are arranged. Using the MaNDi and IMAGINE instruments, the researchers will be able to piece together the protease’s atomic structure by using neutrons to track the hydrogen atoms within the crystallized protein samples.
But first, they have to obtain crystals of high quality that are large enough for neutron experiments. This is where the team has made significant strides early on.
Crystal quality is first determined by how well they diffract, or scatter, x-rays. Typically, this process is conducted at a synchrotron facility, where the crystals might be frozen to around 100 K (or about -280°F).
The team used the Protein Crystallization and Characterization lab at SNS to grow SARS-CoV-2 protease crystals, which took about a week to 10 days. To analyze the quality of the crystals, they used the local x-ray machine, a Rigaku HighFlux HomeLab, which provided several key findings.
First, the x-ray experiments confirmed the crystals were of high quality and that the method used to grow them might produce larger crystals suitable for neutron experiments. Second, having a local machine allowed them to collect x-ray measurements at room temperature, around 70°F.
The room-temperature measurements enabled them to observe the plasticity, or flexibility, of the protease structure, providing discernable information about how the structure behaves in conditions close to the virus’s physiological environment. Those data could not have been obtained using frozen samples.
“This is an important milestone in our effort to do neutron diffraction. The investment in a local x-ray machine has paid off quite well,” said Kneller. “In one instance, we grew crystals on Monday and collected data on them on Tuesday. Otherwise, to obtain that information you would have to send your crystals to a synchrotron, which could take days to weeks.”
“And right now, because of the pandemic, you can’t go to a synchrotron,” added Kovalevsky. “And to analyze crystals at room temperature, you have to be there.”
“The information we learned from the room-temperature structure has the ability to immediately impact the computational directions researchers are using. We found some differences between our room-temperature near-physiological structure and the frozen structures from the synchrotrons, which may be important for the computational work, such as the small-molecule docking studies being done on ORNL’s supercomputer Summit,” said Kneller.
“So far, we’ve been very successful in our early studies of COVID-19. We’ve already submitted a manuscript for publication about our structural findings, in which we’ve essentially conducted two months of research that normally might have taken a year.”
Aiding Kovalevsky and Kneller in the data and structure analysis of the protein crystals was Leighton Coates, an instrument scientist on the SNS MaNDi diffractometer who is also a member of the crystallographic team studying the SARS-CoV-2 protease.
The data generated over the next several months will be shared with other national laboratories, universities, and the broader science community to build more accurate models for computational simulations used to identify potential drug candidates to stop the virus.
“The scientific community has responded swiftly to the COVID-19 pandemic. We are fortunate to be able to make our own contributions by leveraging years of experience studying HIV to build a better understanding of how the novel coronavirus replicates and how we can battle it by inhibiting its essential protease,” said Kovalevsky.
Before the pandemic turned their attention and efforts to researching SARS-CoV-2, Kovalevsky and Kneller had a clear plan for attacking HIV.
Thirty-nine million people around the world are infected with HIV. Providing these people with better treatment options would not only improve their quality of life but also prevent this disease from spreading further.
The HIV protease works by cleaving harmless, or nonfunctional, strands of proteins into smaller proteins, turning them into functional viral proteins that enable the virus to assemble and continue infecting healthy human cells. In general, HIV protease inhibitors are quite effective at blocking protease during HIV replication, but some variations of protease have developed an ability to resist drug inhibitors.
“If we can learn more about the molecular mechanisms that make HIV protease variants drug resistant, we can design drugs that are better equipped to outsmart its defenses,” said Kneller.
Specifically, Kneller and Kovalevsky wanted to explore PRS-17, a unique HIV protease variant that is 10,000 times less likely than other nonresistant variants to be inhibited by the most effective clinical HIV protease inhibitors currently available. Kovalevsky explained that while HIV treatment programs have come a long way since the HIV pandemic first began in the 1980s, mutant variants like PRS-17, resulting from prolonged treatment, could compromise years of pharmaceutical innovation and progress and result in failed antiviral therapies.
“Drug resistance is now the biggest problem for HIV patients. With proper treatment, patients can live long and happy lives with undetectable levels of HIV in their system. They won’t develop AIDS or spread HIV to others. But PRS-17 and other drug-resistant HIV protease variants make it difficult for physicians to combat HIV in their patients,” said Kovalevsky.
Understanding exactly how PRS-17 neutralizes the efficacy of HIV protease inhibitors is difficult, say the researchers. Viruses’ constituent proteins are complex systems, and PRS-17 has the ability to employ several different mechanisms to guard itself against anti-retroviral drugs.
“Figuring out how PRS-17 resists HIV protease inhibitors is a challenge, but one that we absolutely have to overcome. PRS-17 is a clinical isolate, which means it came from an actual patient struggling to combat this disease,” explained Kneller. “Learning more about it could save the lives of many patients, because the knowledge we gain using neutrons on PRS-17 will be transferrable to other similar extremely drug-resistant protease variants.”
The team intended to create a map of the PRS-17 protease to better understand the molecular mechanisms behind its drug resistance. That involved using the MaNDi and VISION instruments at SNS and the IMAGINE instrument at HFIR.
“It was very much the same approach we are now trying with COVID-19,” said Kovalevsky.
With MaNDi and IMAGINE, Kneller and Kovalevsky were planning to probe crystallized samples of PRS-17 protease to generate detailed data on its static atomic structure. Using VISION would enable them to probe powdered samples of PRS-17 protease to provide insights into its dynamic properties by measuring the molecular vibrations.
Neutrons are particularly well-suited to study components of viruses such as HIV (or SARS-CoV-2) because of their sensitivity to hydrogen, an important component of all proteins. With neutron crystallography, the team could precisely locate each hydrogen atom within PRS-17’s protease, giving them unprecedented insight into how the protein functions and what interactions it undergoes with a protease inhibitor.
“Use neutron crystallography at MaNDi and IMAGINE to locate hydrogen atoms in crystals of PRS-17 protease, would enable us to build a comprehensive profile of its static structure,” said Kneller. “With VISION, we would also track hydrogen atoms, but we would use powdered samples of PRS-17 protease that have been rehydrated to mimic the crowded conditions of an HIV viral particle. That would allow us to see its dynamic properties and learn more about how it might move when it is working within a viral particle.”
Kneller explained that getting information about both the static and dynamic properties of PRS-17 is important for developing a complete understanding of this virus’s resistance to anti-retroviral drugs.
“If I tracked your location just once a day at midnight, I would think you spend all of your time at home. But really, you move around quite a bit throughout the day. That’s why it’s important to collect both static and dynamic measurements of our sample. It lets us build a fuller picture of protease’s behavior,” said Kneller.
“Without neutron crystallography, researchers have to make educated guesses about where hydrogen atoms are in a protein whenever they attempt to understand how the protein does its job,” added Kneller. “These types of experiments that Andrey has done previously have actually been able to confirm the locations of these hydrogen atoms in nonresistant HIV protease variants, but never in an extremely drug-resistant protease variant. That means we would be able to produce truly unique and novel data about this protease.”
Kneller and Kovalevsky hope to one day generate data through their experiments that will become an invaluable resource for researchers looking to combat drug-resistant strains of HIV.
“It’s a team effort. Chemists, biologists, and professionals from the pharmaceutical industry all have to work together to combat illness,” said Kneller. “Together, we can develop effective treatments for drug-resistant strains of HIV.”
Research was supported by the DOE Office of Science through the National Virtual Biotechnology Laboratory, a consortium of DOE national laboratories focused on response to COVID-19, with funding provided by the Coronavirus CARES Act.
Source: Oak Ridge National Laboratory
Be the first to comment on "History of Insightful HIV Research Inspires Neutron-Scattering Approach to Studying COVID-19"