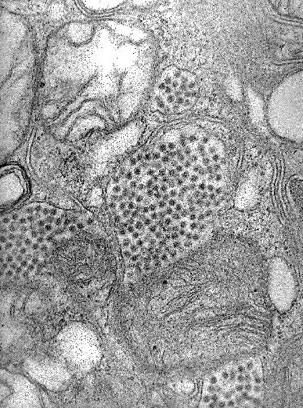
A new study led by researchers at Harvard Medical School has identified a set of cellular receptors for at least three related alphaviruses shared across mosquitoes, humans, and animals that host the virus.
Going a step further, the researchers tested a “decoy” molecule that successfully prevented infection and slowed disease progression in a series of experiments in cells and animal models, an important first step toward developing preventive and curative medicines against these highly pathogenic viruses with pandemic potential.
The results were published December 20 in Nature.
Understanding the basic biology of a virus’s life cycle is crucial to finding a way to prevent an illness, and building such foundational knowledge before an outbreak is essential for preparing for future outbreaks, said study senior author Jonathan Abraham, assistant professor of microbiology in the Blavatnik Institute at HMS and an infectious disease specialist at Brigham and Women’s Hospital.
“Understanding how a virus enters and infects a cell is as basic as it gets,” he said. “Viral entry into human or other mammalian cells marks the beginning of the infection and eventually disease, and is a great place to begin looking for potential preventive strategies and curative medications.”
The alphaviruses the researchers studied, including EEEV, have a history of causing deadly, if short-lived, outbreaks, but little is known about how the virus attacks host cells. Only a few other receptors related to infection from alphaviruses have been identified. This gap in knowledge is one of the reasons for the lack of targeted treatments for these lethal viruses, Abraham said.
Eastern equine encephalitis (triple E) typically spreads to humans from the bite of an infected mosquito. The most recent outbreak of triple E took place in New England in 2019. The triple E outbreak hospitalized all 32 known cases and killed 12 of those infected, according to a U.S. Centers for Disease Control and Prevention report on the 2019 triple E outbreak.
The virus has a 30 percent fatality rate—meaning it kills nearly one-third of those diagnosed with the disease—comparable to Ebola virus disease or smallpox. Nearly half of those who do survive experience long-term neurologic complications from the disease, according to the CDC. Larger outbreaks took place in the 1930s and 1950s, but data collection and diagnostics have changed so much since then that it’s difficult to compare the magnitude of the outbreaks, the CDC report noted.
There are great benefits to doing this work before the onset of major outbreaks, said Abraham, who has worked on identifying receptors that viruses use as pathways for gaining entry into cells and causing disease and on developing antibody treatments to prevent SARS-CoV-2 infection and COVID-19.
For example, prior work on SARS-CoV during the SARS outbreak of the early 2000s was crucial to increasing preparedness against SARS-CoV-2. The availability of SARS-CoV-2 genome sequences within days of the initial reporting of the novel virus were crucial to the rapid development of vaccines and antibody treatments for COVID-19, since the closely related viruses that cause COVID-19 and SARS both attack human hosts by using the same receptor on human cells to gain entry and cause disease.
New screening tools and techniques in molecular biology, protein biochemistry, biophysics, and structural biology provide unprecedented power to learn more than ever about the basic biology of viruses before they emerge as global threats, Abraham said.
"The time to prepare for these uncertain but potentially catastrophic scenarios is not when they occur but well before they do,” Abraham said.
For the current study, the researchers first used a CRISPR-Cas9 gene-editing screen to identify a receptor for Semliki Forest virus (SFV) on human cells. SFV is an alphavirus that can cause severe neurologic disease and death in rodents and other animals.
The receptors that the researchers found for SFV were also compatible with EEEV and another related virus called Sindbis, which can cause fever and severe joint pain in humans and causes neurological disease in animals and rodents
“That’s why it’s important to study these viruses as families,” Abraham said. “You can end up studying a virus like SFV and discover something really exciting about the biology of related viruses that has the potential to unlock novel ways to treat new categories of viruses that are capable of causing serious disease and outbreaks in humans.”
Identifying a receptor for multiple viruses would give the scientists and physicians a running start on developing tools to prevent, control, and treat infections should an outbreak of one of the viruses occur, Abraham said.
As a means of verifying that the receptors in question were important in causing infection, the researchers conducted experiments with a decoy protein, a molecule with a structure that mimics the receptor and can trick the virus into binding to the drug instead of to the host cell it aims to infect. The molecule, in effect, disables the virus and averts entry into the host cell, preventing infection.
The team’s experiments demonstrated that blocking the virus from interacting with the host cell receptor prevented infection of human and mouse neurons. They also found that the decoy molecule protected infected mice from developing rapidly fatal alphavirus encephalitis—a finding that the researchers say suggests this pathway could be targeted by drugs or antibodies to treat alphavirus encephalitis in humans when infections do occur. The researchers caution that they conducted their animal studies with Semliki Forest virus and not with EEEV, so further experiments would be required to verify that the same approach could work for different alphaviruses and in humans.
Turning a basic insight like this into a clinical tool usually takes years. Researchers need to make sure it is safe and effective and need to find the best way to administer the decoy molecules. To buy as much time in advance to prepare for emerging viruses, it is critical to build this base of knowledge before the next pandemic, Abraham said.
“All of us—scientists, policymakers, and citizens—would benefit greatly from learning to be more forward thinking,” Abraham said. “The more we know about the basic biology of different families of viruses, especially how they infect and interact with their hosts, the better prepared we will be to face whatever comes next.”
Co-authors included Lars Clark, Sarah Clark, ChieYu Lin, Adrian Coscia, Katherine Nabel, Dylan Neel, Isaac Chiu, Paula Montero Llopis, Tracy Young-Pearse, Pan Yang, and Vesna Brusic at Harvard Medical School; Asim Ahmed at Boston Children's Hospital; Jianying Liu and Kenneth Plante, and Scott Weaver at the University of Texas Medical Branch; and Iryna Stryapunina and Flaminia Catteruccia at the Harvard T.H. Chan School of Public Health.
This research was supported by the U.S. Department of Health & Human Services, the National Institutes of Health, the Burroughs Wellcome Fund, the William Randolph Hearst Foundation, the Brigham and Women's Hospital Harvard Milton Fund , the Vallee Scholar Award, and the Howard Hughes Medical Institute.
DOI
https://doi.org/10.1038/s41586-021-04326-0