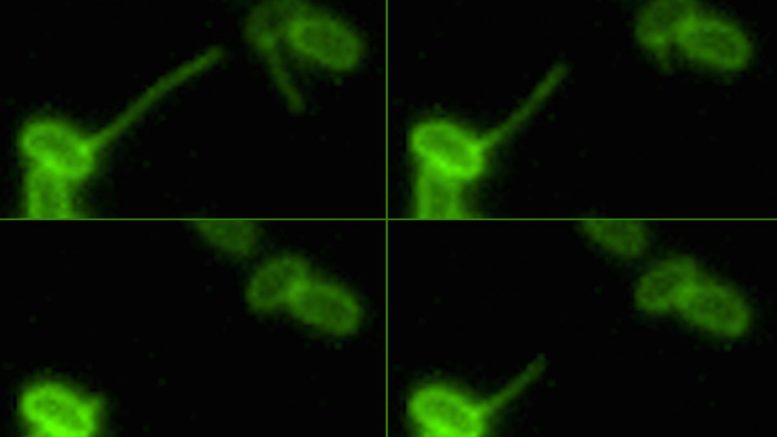
A bacterium retracts its pili, reeling in a piece of DNA in the environment. This action facilitates "natural transformation," a process by which bacterium acquire new genetic traits, including antibiotic resistance. Courtesy the Dalia Lab, Indiana University
A new study from Indiana University has revealed a previously unknown role a protein plays in helping bacteria reel in DNA in their environment -- like a fisherman pulling up a catch from the ocean.
The discovery was made possible by a new imaging method invented at IU that let scientists see for the first time how bacteria use their long and mobile appendages -- called pili -- to bind to, or "harpoon," DNA in the environment. The new study, reported Oct. 18 in the journal PLOS Genetics, focuses on how they reel their catch back in.
By revealing the mechanisms involved in this process, the study's authors said the results may help hasten work on new ways to stop bacterial infection.
"The issue of antibiotic resistance is very relevant to this work since the ability of pili to bind to, and 'reel in,' DNA is one of the major ways that bacteria evolve to thwart existing drugs," said Ankur Dalia, an assistant professor in the IU Bloomington College of Arts and Sciences' Department of Biology, who is senior author on the study. "An improved understanding of this 'reeling' activity can help inform strategies to stop it."
The act of gobbling up and incorporating genetic material from the environment -- known as natural transformation -- is an evolutionary process by which bacteria incorporate specific traits from other microorganisms, including genes that convey antibiotic resistance.
The need for new methods to stop bacterial infection is growing since overuse of existing antibiotics, which speeds how quickly infectious organisms evolve to outsmart these drugs, is causing the world to quickly run out of effective treatments. By 2050, it's estimated that 10 million people could die each year from antimicrobial resistance.
Although they may look like tiny arms under a microscope, Dalia said, pili are actually more akin to an erector set that is quickly put together and torn down over and over again. Each "piece" in the structure is a protein sub-unit called the major pilin that assembles into a filament called the pilus fiber.
"There are two main motors that had previously been implicated in this polymerization and depolymerization process," added Jennifer Chlebek, a Ph.D. student in Dalia's lab, who led the study. "In this study, we show that there is a third motor involved in the depolymerization process, and we start to unravel how it works."
The two previously characterized "motors" that control the pili's activity are the proteins PilB, which constructs the pili, and PilT, which deconstructs it. These motors run by utilizing ATP, a source of cellular energy. In this study, IU researchers showed that stopping this process, which switches off the power to PilT, does not prevent the retraction of the pili, as previously thought.
Instead, they found that a third motor protein, called PilU, can power pilus retraction even if PilT is inactive, although this retraction occurs about five times more slowly. The researchers also found that switching off power to both retraction proteins slows the retraction process to a painstaking rate of 50 times slower. An unaltered pilus retracts at a rate of one-fifth of a micron per second.
Moreover, the study found that switching off PilU affects the strength of pilus retraction, which was measured by collaborators at Brooklyn College. The study also showed that PilU and PilT do not form a "hybrid" motor, but instead that these two independent motors somehow coordinate with one another to mediate pilus retraction.
"While the PilU protein had previously been implicated in pilus activity, its exact role has been difficult to determine because cells that lack this protein generally only have very subtle effects," Chlebek added. "Our observation that PilU can support pilus retraction in a mutant strain, when we threw a wrench in the PilT motor, was the key to unlocking how this protein aids in the depolymerization of pili."
The ability to precisely measure the pili's retraction rate -- and therefore precisely measure the impact of altering the proteins that affect this process -- was made possible by the ability to see pili under a microscope, which was not possible until the breakthrough imaging method invented at IU.
"The ability to fluorescently dye the pili was huge," Dalia said. "It allowed us to not only see the pili's activity but also measure it in ways which simply would not have been possible in the past."
Next, Chlebek aims to learn more about how the pili still retract when the power is switched off to both retraction motors, as well as explore how these insights could apply to understanding pili activity in other strains of bacteria.
This research was supported in part by the National Institutes of Health. Additional IU authors on the study were Brittany Herrin and Hannah Hughes, both Ph.D. students; Triana Dalia, a research associate in the Department of Biology; and Joseph Che-Yen Wang, an assistant scientist at the IU Electron Microscopy Center. The researchers at Brooklyn College were Aleksandra S. Ratkiewicz, Rasman Rayyan and Nicolas Biais.
Be the first to comment on "DNA-Reeling Bacteria Yield New Insight on How Superbugs Acquire Drug Resistance"