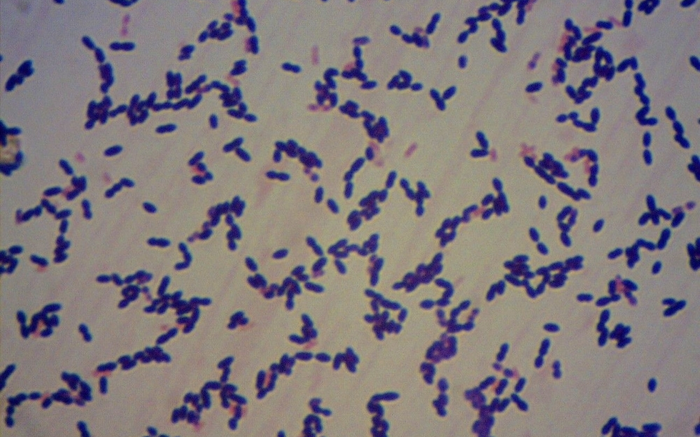
A study published in the journal PLOS ONE could one day help healthcare professionals determine whether bacteria of the species Streptococcus pneumoniae, which cause meningitis are resistant to antibiotics.
This type of analysis is no easy task when the conventional method is used. The bacteria must be isolated from a patient sample and analyzed while still alive, which is difficult because the microorganisms are sensitive and usually do not survive the journey to the laboratory.
A highly feasible novel method has been developed in Brazil by researchers at the Santo André branch of Adolfo Lutz Institute (IAL), São Paulo state's central epidemiological surveillance laboratory. Between 2014 and 2020, they analyzed 873 samples of cerebrospinal fluid from patients with suspected streptococcal meningitis at health clinics in six cities in the state – Diadema, Mauá, Santo André, São Bernardo do Campo, São Caetano do Sul and Ribeirão Pires. Cerebrospinal fluid is produced by tissue that lines the ventricles in the brain. It flows in and around the brain and spinal cord to cushion them from injury and provide nutrients.
As part of the laboratory’s routine, the scientists analyzed the samples using real-time PCR, the gold standard for diagnosing infectious diseases, including COVID-19. The technique amplifies a specific gene or gene sequence from the target microorganism, if present in the sample, so that it can be identified more easily. In this case, S. pneumoniae (pneumococcus) was detected in 149 samples.
They then re-analyzed the samples that tested positive for pneumococcus to detect the three genes associated with resistance to antibiotics, again using real-time PCR but this time with SYBR Green, a dye that binds to DNA and emits a fluorescent signal which is captured by the equipment.
To find out which classes of antibiotics the bacteria resisted – penicillin, lincosamides or macrolides – they used the dissociation curve technique. “This technique entails raising the temperature of the samples degree by degree, making the dye separate from the DNA as the twin strands in the double helix forming the genetic material amplified in the PCR machine gradually unwind. We measured the melting temperature [Tm], which is when half the structure is still entwined and the rest has separated out. This temperature varies depending on the amplified gene, so it can be used to identify the gene that’s been amplified and hence the antibiotic to which the bacteria are resistant,” said Ivana Campos, principal investigator for the study.
After conducting all these procedures, the researchers compared the results with those obtained by the conventional method used to analyze resistance to antibiotics, in which live bacteria are observed while in contact with each drug to see if they are able to proliferate. This conventional test was performed on 25 samples, which were the only ones that contained viable pneumococci for the procedure. The results were similar, confirming the novel technique’s potential.
“We found that 51% of the samples analyzed, which IAL received between 2014 and 2020, were sensitive to antibiotics. That’s positive, meaning these patients must have had a good prognosis. On the other hand, 17% were resistant to various drugs, which is very dangerous because in these cases it’s more difficult to treat the disease and other classes of antibiotic have to be tried,” Campos said.
Moreover, S. pneumoniae is capable of changing its genetic makeup as it reproduces, so that new copies have the genes associated with drug resistance. “We, therefore, concluded that the test we developed can be used to study the resistance profile of pneumococcus even in the absence of the isolated strains, as evidenced for our region,” she said.
Source: São Paulo Research Foundation (FAPESP)