Scientists have discovered why some coronaviruses are more likely to cause severe disease, which has remained a mystery, until now. Researchers of the University of Bristol-led study, published in Science Advances today, say their findings could lead to the development of a pan-coronavirus treatment to defeat all coronaviruses—from the 2002 SARS-CoV outbreak to Omicron, the current variant of SARS-CoV-2, as well as dangerous variants that may emerge in future.
In this new study, an international team, led by Bristol’s professor Christiane Schaffitzel, scrutinized the spike glycoproteins decorating all coronaviruses. They reveal that a tailor-made pocket feature in the SARS-CoV-2 spike protein, first discovered in 2020, is present in all deadly coronaviruses, including MERS and Omicron. In striking contrast, the pocket feature is not present in coronaviruses which cause mild infection with cold-like symptoms.
The team members say their findings suggest that the pocket, which binds a small molecule, linoleic acid—an essential fatty acid indispensable for many cellular functions including inflammation and maintaining cell membranes in the lungs so that we can breathe properly—could now be exploited to treat all deadly coronaviruses, at the same time rendering them vulnerable to a linoleic acid-based treatment targeting this pocket.
COVID-19, caused by SARS-CoV-2, is the third deadliest coronavirus outbreak following SARS-CoV in 2002 and MERS-CoV in 2012. The much more infectious SARS-CoV-2 continues to infect people and damage communities and economies worldwide, with new variants of concern emerging successively, and Omicron evading vaccination and immune response.
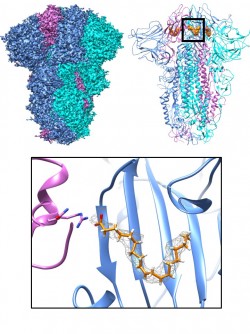
Schaffitzel, from Bristol’s School of Biochemistry, explained, “In our earlier work we identified the presence of a small molecule, linoleic acid, buried in a tailor-made pocket within the SARS-Cov-2 glycoprotein, known as the ‘Spike protein’, which binds to the human cell surface, allowing the virus to penetrate the cells and start replicating, causing widespread damage. We showed that binding linoleic acid in the pocket could stop virus infectivity, suggesting an anti-viral treatment. This was in the original Wuhan strain that started the pandemic. Since then, a whole range of dangerous SARS-CoV-2 variants have emerged, including Omicron, the currently dominating variant of concern. We scrutinised every new variant of concern and asked whether the pocket function is still present.”
Omicron has undergone many mutations, enabling it to escape immune protection offered by vaccination or antibody treatments that lag behind this rapidly evolving virus. Intriguingly, while everything else may have changed, the researchers found that the pocket remained virtually unaltered, also in Omicron.
Christine Toelzer, research associate in the School of Biochemistry and lead author of the study, added, “When we realized that the pocket we had discovered remained unchanged, we looked back and asked whether SARS-CoV and MERS-CoV, two other deadly coronaviruses causing previous outbreaks years ago, also contained this linoleic acid binding pocket feature.”
The team applied high-resolution electron cryo-microscopy, cutting-edge computational approaches and cloud computing. Their results showed that SARS-CoV and MERS-CoV also had the pocket, and could bind the ligand, linoleic acid, by a virtually identical mechanism.
Schaffitzel concluded, “In our current study, we provide evidence that the pocket remained the same in all deadly coronaviruses, from the first SARS-CoV outbreak 20 years ago to Omicron today. We have shown previously that linoleic acid binding to this pocket induces a locked spike, abrogating viral infectivity. We also show now that linoleic acid supplementation suppresses virus replication inside cells. We anticipate that future variants will also contain the pocket, which we can exploit to defeat the virus.”
Reference: “The free fatty acid-binding pocket is a conserved hallmark in pathogenic b-coronavirus spike proteins from SARS-CoV to Omicron” by C Toelzer et al. in Science Advances
Source: University of Bristol