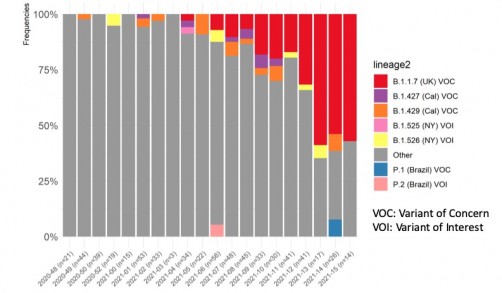
Graph courtesy of UT Southwestern Medical Center
UT Southwestern Medical Center scientists have identified the first cases of the Brazilian variant of COVID-19 infection in North Texas using next-generation sequencing technologies along with PCR testing.
UT Southwestern sequencing protocols show the U.K. variant (B.1.1.7) has become dominant in North Texas – found in about 55% of sampled individuals – followed by the California (B.1.429), New York (B.1.526), and Brazil variants (P.1), which were each found in about 7% of those sampled. For the last several months, all positive COVID-19 cases at UT Southwestern have been tested for variants – currently about 20-30 samples per week. Although a relatively small sample, ongoing sequencing gives a better picture of how frequent the variants are, and whether the prevalence of the Brazil variant, for example, is gaining traction in Texas. UT Southwestern reported the finding to Dallas County Health and Human Services (DCHHS) on April 24, 2021.
The emergence of the Brazil variant in North Texas is considered worrisome, listed as a Variant of Concern by the Centers for Disease Control and Prevention because it is more easily transmitted and is less susceptible to antibodies. Antibody treatments (convalescent plasma or monoclonal antibodies) and post-vaccine antibodies have a more difficult time binding to the Brazil variant of the SARS-CoV-2 virus that causes COVID-19. This can render vaccines less effective, a factor known as vaccine escape.
“These findings reinforce the importance of vaccination – which helps slow the transmission of all types of virus and protects against more severe disease – and continuing to follow safeguards such as masking and distancing,” says Jeffrey SoRelle, MD, assistant instructor of pathology. “Even though the variant is known to evade the immune system, meaning less vaccine efficacy, vaccine protection is far better than having none.”
In particular, the vaccines appear to provide protection against severe disease and death, underscoring the urgency of vaccination efforts. The presence of the Brazil variant also reinforces why social distancing and masking continue to be important: To provide additional protection in addition to vaccine if or when it is less effective against a variant.
Because each variant has different rates of transmissibility, identifying the different variants appearing in North Texas is important for accurate modeling to predict prevalence, which in turn helps health care officials better prepare for surges, increased hospitalizations, and needed resources. Prevalence of variants can also influence treatment.
“An important part of forecasting is predicting how quickly the disease will spread, so knowing which variants are prevalent helps us make more accurate models,” says SoRelle, whose research is part of Genomics and Molecular Pathology. Transmissibility rates of the variants ranges from 20% to more than 50%. The transmissibility of the Brazil variant is currently not well documented, but may be up to two times more transmissible.
The sequencing was performed in the McDermott Next-Generation Sequencing Core under the supervision of Helen H. Hobbs, MD, a Howard Hughes Medical Institute Investigator and Professor of Internal Medicine and Molecular Genetics who directs the McDermott Center for Human Growth and Development at UT Southwestern. SoRelle is collaborating with Hobbs by providing all positive COVID-19 samples tested at UT Southwestern and corroborating results with a rapid, focused PCR- based test. The collaboration with the McDermott Center Next Generation Sequencing (NGS) Core at UT Southwestern, allows whole genome sequencing of the SARS-CoV-2 virus in a state-of-the-art facility that performs NGS coupled with bioinformatic analysis. Whole genome sequencing is the best way to monitor for any new variants. Variant data will be integrated into forecasting models for epidemiologic predictions.
Source: UT Southwestern Medical Center
889967 7215I was suggested this weblog by way of my cousin. Im no longer positive whether or not this put up is written by him as nobody else realize such detailed about my trouble. You are great! Thanks! 416908